A team of researchers led by Feng Zhang at the Broad Institute of MIT and Harvard and the McGovern Institute for Brain Research at MIT has discovered the first programmable RNA-guided system in eukaryotes—organisms that include fungi, plants, and animal
In a study published in Nature, the team describes how the system is based on a protein called Fanzor. They showed that Fanzor proteins use RNA as a guide to precisely target DNA, and that Fanzors can be reprogrammed to edit the genome of human cells. Compact Fanzor systems have the potential to be more easily delivered to cells and tissues as therapeutics than CRISPR/Cas systems, and further refinements to improve their targeting efficiency could be an important new technology for editing the human genome.
CRISPR/Cas was first discovered in prokaryotes (bacteria and other single-cell organisms that lack nuclei) and scientists including Zhang’s lab have long wondered whether similar systems exist in eukaryotes. The new study shows that RNA-guided DNA-cutting mechanisms are present in all kingdoms of life.
“CRISPR-based systems are widely used and powerful because they can be easily reprogrammed to target different sites in the genome,” said Zhang, senior author on the study and a member of the core institute at Broad, an investigator at MIT’s McGovern Institute, the James and Patricia Poitras Professor of Neuroscience at MIT, and a Howard Hughes Medical Institute investigator. “This new system is another way to make precise changes in human cells, complementing the genome editing tools we already have.”
Exploring the domains of life
The main goal of the Zhang lab is to develop genetic medicines using systems that can modulate human cells by targeting specific genes and processes. “A few years ago, we started asking, ‘What is beyond CRISPR, and are there other RNA-programmable systems in nature?'” Zhang said.
Two years ago, members of the Zhang lab discovered a class of RNA-programmable systems in prokaryotes called OMEGA, which are often associated with transposable elements, or “jumping genes,” in bacterial genomes and probably gave rise to the CRISPR/Cas system. That work also highlighted similarities between prokaryotic OMEGA systems and Fanzor proteins in eukaryotes, suggesting that Fanzor enzymes may also use an RNA-guided mechanism to target and cut DNA.
In the new study, the researchers continued their study of the RNA-guided system by isolating Fanzors from fungi, algae, and amoeba species, in addition to a clam known as the Northern Quahog. Co-first author Makoto Saito of the Zhang lab led the biochemical characterization of Fanzor proteins, showing that they are DNA-cutting endonuclease enzymes that use nearby non-coding RNAs known as ωRNAs to target specific sites in the genome. This is the first time this mechanism has been found in eukaryotes, such as animals.
Unlike CRISPR proteins, Fanzor enzymes are encoded in eukaryotic genomes within transposable elements and the team’s phylogenetic analysis suggests that Fanzor genes moved from bacteria to eukaryotes through called horizontal gene transfer.
“These OMEGA systems are more ancestral to CRISPR and they are among the most abundant proteins on the planet, so it makes sense that they were able to switch back and forth between prokaryotes and eukaryotes,” Saito said.
To explore Fanzor’s potential as a genome editing tool, the researchers demonstrated that it can generate insertions and deletions at targeted genome sites within human cells.
The researchers discovered that the Fanzor system was initially less efficient at snipping DNA than CRISPR/Cas systems, but through systematic engineering, they introduced a combination of protein mutations that increased its activity by 10-fold. Additionally, unlike some CRISPR systems and the OMEGA protein TnpB, the team found that a fungal-derived Fanzor protein did not exhibit “collateral activity,” in which an RNA-guided enzyme removes the target of Its DNA as well as degrades nearby DNA or RNA. The results suggest that Fanzors can be developed as efficient genome editors.
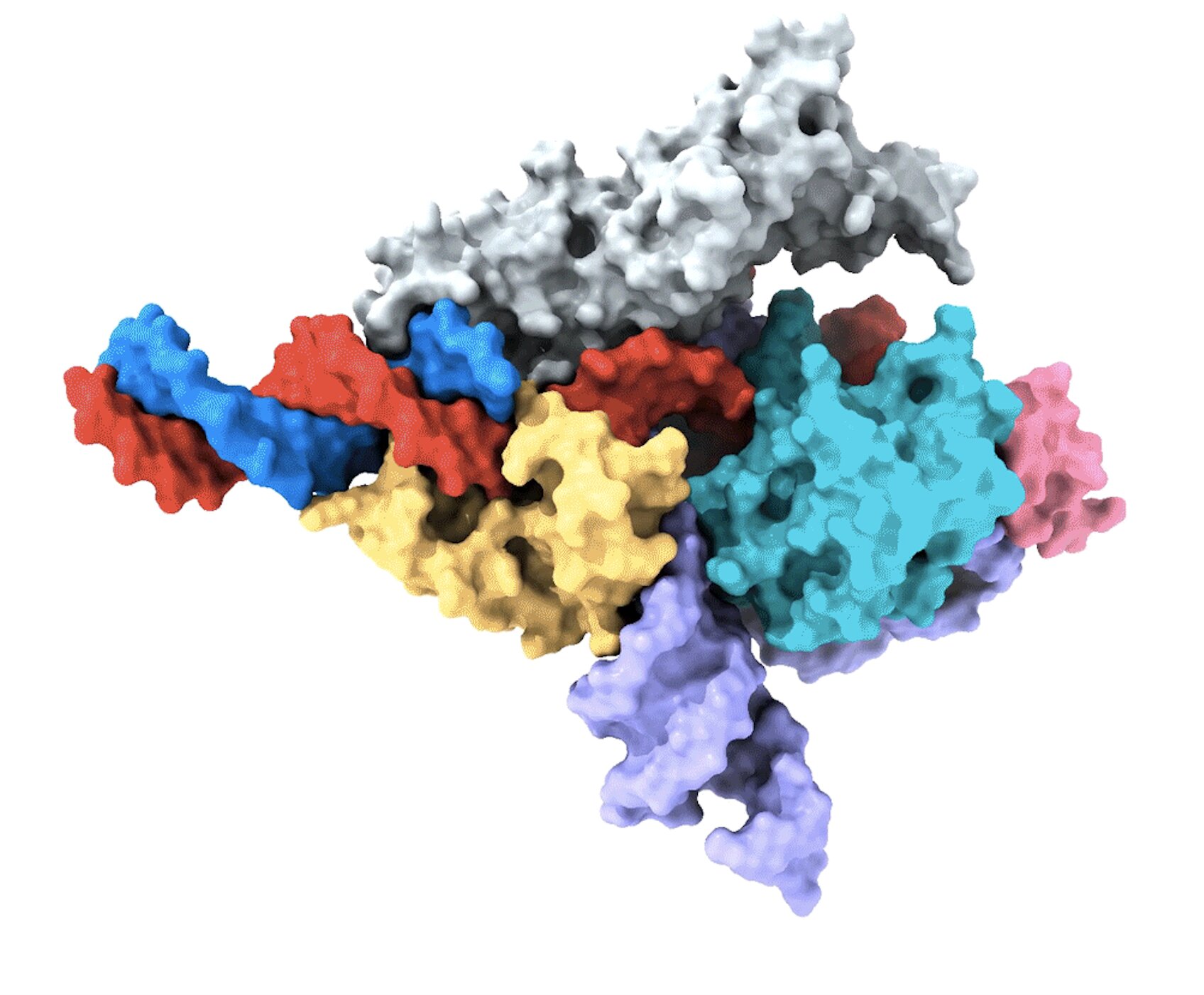
Co-first author Peiyu Xu led the effort to study the molecular structure of the Fanzor/ωRNA complex and describe how it binds to DNA to cut it. Fanzor shares structural similarity with its prokaryotic counterpart CRISPR-Cas12 protein, but the interaction between ωRNA and the catalytic domains of Fanzor is more extensive, suggesting that ωRNA may play a role in catalytic reactions. “We are excited about these structural insights for helping us further engineer and optimize Fanzor for improved efficiency and accuracy as a genome editor,” said Xu.
Like CRISPR-based systems, the Fanzor system can easily be reprogrammed to target specific genome sites, and Zhang says it could one day develop into a powerful new genome-editing technology for research and therapeutic applications. The abundance of RNA-guided endonucleases such as Fanzors further expands the number of OMEGA systems known across the kingdoms of life and suggests that there are more to be discovered.
“Nature is amazing. There is so much diversity,” Zhang said. “There are probably more RNA-programmable systems out there, and we’re continuing to explore and hopefully discover more.”
Additional information:
Fanzor is a eukaryotic programmable RNA-guided endonuclease, Nature (2023). DOI: 10.1038/s41586-023-06356-2
Journal information:
Nature
#Researchers #discovered #CRISPRlike #system #animals #edit #human #genome














Add Comment